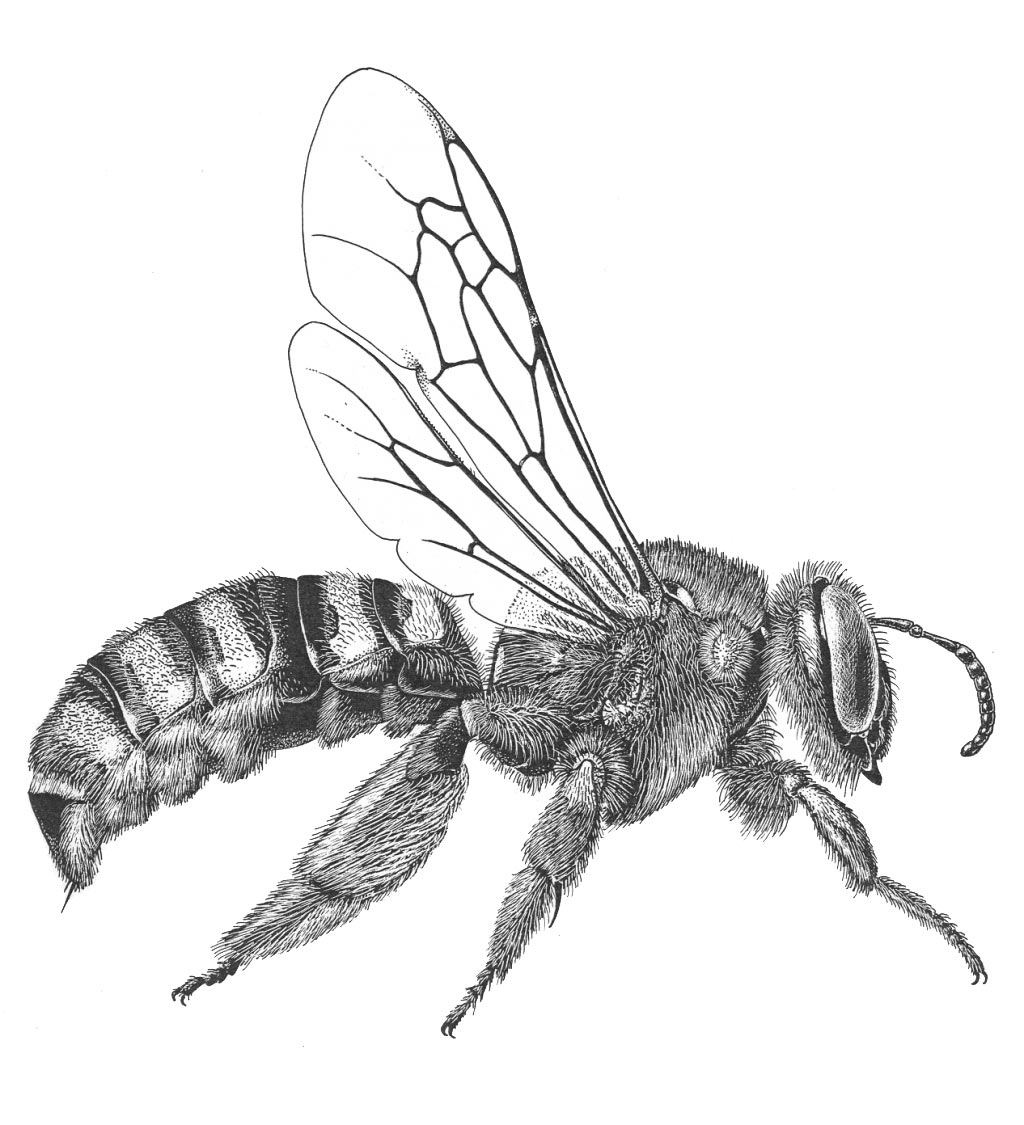
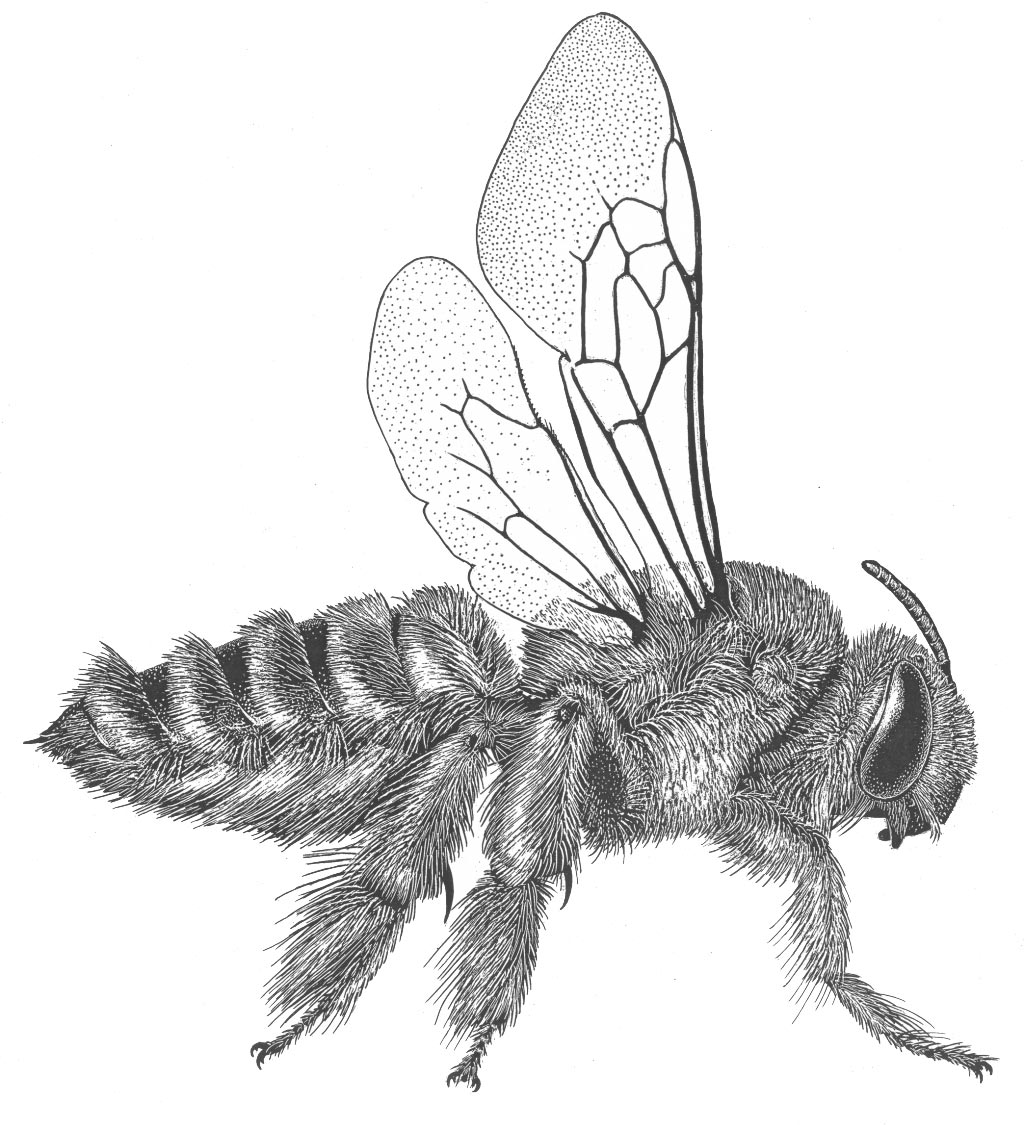
One major focus of the lab is understanding the phylogeny and evolutionary history of bees and related wasps (Aculeata).
We use cutting-edge methods to develop large molecular data sets for resolving phylogenetic relationships among bees at various taxonomic levels. The lab has recently embarked on a new collaborative project on the higher-level phylogeny of bees using a combination of ultra-conserved elements (UCEs) and low-coverage genome (LCG) sequencing. This project involves a collaboration with Elizabeth Murray (Washington State University), Silas Bossert (Washington State University), Michael Branstetter (USDA-ARS Pollinating Insects Research Unit) and Paulmichael Maxfield (Natural History Museum of Utah). The project, entitled “Bees of the World – Phylogenomics, Biogeography, and Evolution of Host-Plant Associations” (DEB-2127744), will allow us to resolve many of the outstanding issues in bee phylogeny, historical biogeography, and host-plant evolution.
Current Projects
The interaction between bees and flowering plants represents one of the most important co-evolutionary stories on earth. The origin of bees (between 110 and 130 my ago) coincides with a rapid increase in angiosperm diversification and the appearance of the largest and most species-rich clade of flowering plants – the eudicots. Flowering plants, with over 350,000 described species, are currently the dominant group of plants in all terrestrial habitats and their dominance is due, at least in part, to their partnership with mobile pollinators – primarily insects and most importantly, bees. Bees provide angiosperm plants with pollination services and plants, in return, provide bees with food for building nests, attracting mates, and rearing offspring. One key to unraveling how the bee-angiosperm partnership evolved over time is to reconstruct both the temporal and spatial patterns of bee diversification. We propose to develop a densely sampled phylogeny of bees at the generic, subgeneric, and species level to examine both historical biogeography as well as patterns of host-plant associations through time.

Previous studies of bee phylogeny have been limited in taxon sampling, geographic sampling, and the number of phylogenetic markers used. The most thorough prior analysis of global bee phylogeny included just 6% of species, 67% of genera, and only a handful of nuclear genes. In order to reconstruct a robust phylogeny of bees, which includes over 20,000 described species, we need more exhaustive taxon sampling across a broader geographic scale, with tools that leverage the power of modern, high throughput phylogenomics.
Our project has three core aims:
Aim 1) Phylogeny — Assemble a comprehensive phylogenomic dataset for bees at the generic level using a combination of low coverage genomes (LCGs) and ultraconserved elements (UCEs).
Aim 2) Historical biogeography — Using the bee phylogeny obtained in Aim 1), compile information on bee fossils and geography to (a) infer a time-calibrated phylogeny for bees and (b) reconstruct historical biogeographic patterns, including vicariance and dispersal.
Aim 3) Host-plant evolution — Using the dated bee phylogeny from Aim 1), analyze patterns of hostplant use in bees in order to (a) reconstruct how host-plant specialization and generalization have evolved over time, (b) test for congruence between bee and angiosperm phylogeny in host-plant specialist lineages of bees, and (c) assess how changes in host-plant breadth have impacted rates of diversification in bees.

In collaboration with colleagues in North America, Europe and Africa, we recently analyzed the phylogenetic relationships among the brood parasitic bee tribes Biastini, Neolarrini, and Townsendiellini. These enigmatic brood parasitic groups form a clearly defined monophyletic group that includes the genus Schwarzia – a rare but fascinating genus of brood parasites from Africa. Our phylogeny is based on analysis of 773 loci generated using probes for ultraconserved elements (UCEs). Our phylogeny provides a basis for analysis of historical biogeography, host-parasite associations, and a revised tribal classification. Lastly, our continued efforts to find the rare Schwarzia in Eastern Africa led to the discovery of three new species, which are described in the paper.